|
To receive announcements of new genome
assembly releases, new software features, updates and
training seminars by email, subscribe to the
genome-announce mailing list.
06 March 2014 -
The new GRCh38 Human Genome Browser is here!
In the final days of 2013, the
Genome Reference Consortium (GRC) released the eagerly awaited GRCh38
human genome assembly, the first major revision of the human genome in more than four
years. During the past two months, the UCSC team has been
hard at work building a browser that will let our users explore the new
assembly using their favorite Genome Browser features and tools.
Today we're announcing the release of a preliminary browser on the GRCh38
assembly. Although we still have plenty of work ahead of us in constructing the rich
feature set that our users have come to expect, this early release will allow you to
take a peek at what's new.
Starting with this release, the UCSC Genome Browser version numbers for human
assemblies will match those of the GRC to minimize version confusion. Hence, the GRCh38
assembly is referred to as hg38 in Genome Browser datasets and documentation. We've
also made some slight changes to our chromosome naming scheme that affect primarily the
names of haplotype chromosomes, unplaced contigs and unlocalized contigs. For more details
about this, see the
hg38 gateway page.
What's new in GRCh38?
-
Alternate sequences - Several human chromosomal regions exhibit sufficient
variability to prevent adequate representation by a single sequence. To address this, the
GRCh38 assembly provides alternate sequence for selected variant regions through the
inclusion of alternate loci scaffolds (or alt loci). Alt loci are
separate accessioned sequences that are aligned to reference chromosomes. This assembly
contains 261 alt loci, many of which are associated with the LRC/KIR area of chr19 and the
MHC region on chr6. (See the
sequences page for a complete list of the
reference chromosomes and alternate sequences in GRCh38.)
-
Centromere representation - Debuting in this release, the large megabase-sized gaps
that were previously used to represent centromeric regions in human assemblies have been
replaced by sequences from centromere models created by
Karen Miga et al., using centromere databases developed during her
work in the Willard lab at
Duke University and analysis software developed while working in the
Kent lab at UCSC.
The models, which provide the
approximate repeat number and order for each centromere, will be useful for read mapping
and variation studies.
-
Mitochondrial genome - The mitochondrial reference sequence included in the GRCh38
assembly and hg38 Genome Browser (termed "chrM" in the browser) is the
Revised Cambridge Reference Sequence (rCRS) from
MITOMAP with GenBank accession
number J01415.2 and RefSeq accession number NC_012920.1. This differs from the chrM
sequence (RefSeq accession number NC_001907) used by the previous hg19 Genome Browser,
which was not updated when the GRCh37 assembly later transitioned to the new version.
-
Sequence updates - Several erroneous bases and misassembled regions in GRCh37 have
been corrected in the GRCh38 assembly, and more than 100 gaps have been filled or reduced.
Much of the data used to improve the reference sequence was obtained from other genome
sequencing and analysis projects, such as the 1000 Genomes Project.
-
Analysis set - The GRCh38 assembly offers an "analysis set" that was
created to accommodate next generation sequencing read alignment pipelines. Several
GRCh38 regions have been eliminated from this set to improve read mapping.
The analysis set may be downloaded from the Genome Browser
downloads page.
For more information about the files included in the GRCh38 GenBank submission,
see the
GRCh38 README. The GRCh38 GenBank record provides a detailed
array of statistics about this assembly.
Bulk downloads of the sequence and annotation data may be obtained from the Genome
Browser FTP server
or the
Downloads
page. The annotation tracks for this browser were generated by UCSC and collaborators
worldwide.
Much more to come! This initial release of the hg38 Genome Browser provides a
rudimentary set of annotations. Many of our annotations rely on data sets from external
contributors (such as our popular SNPs tracks) or require massive computational effort
(our comparative genomics tracks). In the upcoming months/years, we will release many more
annotation tracks as they become available. To stay abreast of new datasets, join our
genome-announce mailing list or follow us on
twitter.
We'd like to thank our GRC and NCBI collaborators who worked closely with
us in producing the hg38 browser. Their quick responses and helpful feedback were a
key factor in expediting this release. The production of the hg38 Genome Browser was
a team effort, but in particular we'd like to acknowledge the engineering efforts of Hiram
Clawson and Brian Raney, the QA work done by Steve Heitner, project guidance
provided by Ann Zweig, Robert Kuhn, and Jim Kent, and documentation work by Donna
Karolchik.
See the Credits page for a detailed
list of the organizations and individuals who contributed to this release.
04 March 2014 -
Introducing new Genome Browser highlight feature
We are excited to announce a new highlight feature in the UCSC
Genome Browser. Using drag-and-select, you can now highlight a
region or gene of interest.
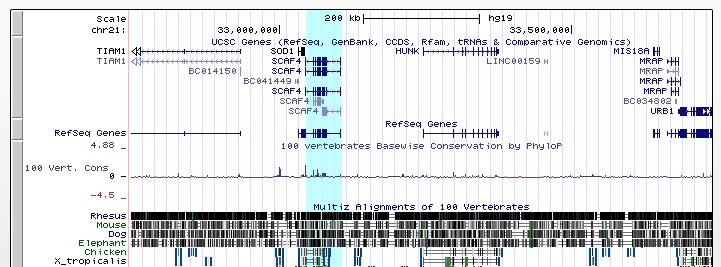
To highlight a region: Click and hold the mouse button on one edge
of the desired area to be highlighted in the Base Position track,
drag the mouse right or left to highlight the selection area, then
release the mouse button. Click the "Highlight" button on the
"drag-and-select" popup. More details about this new feature
can be found on our
help page.
Credit goes to Tim Dreszer, Larry Meyer, Robert Kuhn and Luvina Guruvadoo
for the design, development and testing of this feature. Additional
testing was also provided by several members of the QA team.
28 February 2014 -
New! Expanded onsite workshop program!:
Explore the full power of the UCSC Genome Browser!
Thanks to the funding support of NHGRI, we can now offer hands-on Genome Browser
training onsite at your institution.
Read more.
|
|

